MR design
This study was conducted according to STROBE-MR guidelines11, 12. MR was designed based on three assumptions. (1) Assumption of association: Genetic variations are significantly associated with metabolic-related lifestyle and clinical risk factors. (2) Independence assumption: Genetic variation is independent of other confounders in DSC. (3) Exclusion restriction assumption: Genetic variation influences DSC only through metabolic-related lifestyle and clinical risk factors.13 (Figure 1). Because of the wide variety of metabolic-related lifestyle and clinical risk factors, we have categorized the 19 risk factors into four different groups to aid clarity and understanding. Overall, 19 major metabolic-related lifestyle and clinical risk factors were selected and categorized into four groups: lifestyle factors, physical factors, serum parameters, and metabolic comorbidities. The association of these metabolic-related lifestyle and clinical risk factors with the development of DSC (including EC, GC, CRC, HCC, BTC, and PC) was preliminarily investigated.
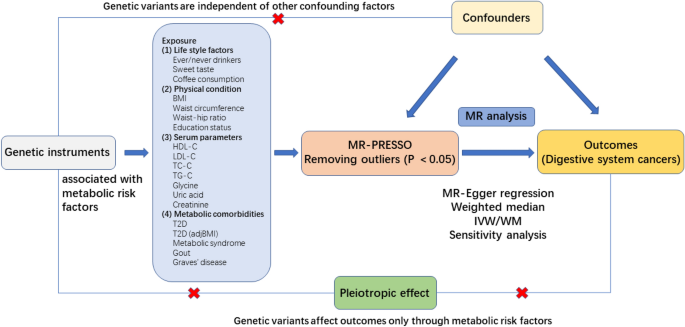
Flowchart of the data collection, processing, and analysis procedures for this study.
Selection of genetic variants
In our MR analysis, we used IV to investigate potential associations between metabolic-related lifestyle and clinical risk factors and the development of DSC. Risk factors/characteristics were classified into four categories. (1) Lifestyle factors, i.e., ever/never drank alcohol, sweet taste, and coffee intake. (2) physical factors, namely body mass index (BMI), waist circumference (adjusted for BMI), waist-hip ratio (adjusted for BMI), and education level; (3) serum parameters, i.e., high-density lipoprotein cholesterol (HDL-C), low-density lipoprotein cholesterol (LDL-C), total cholesterol (TC), triglyceride (TG), glycine, uric acid, and creatinine levels; 4) metabolic comorbidities, namely type 2 diabetes (T2D), T2D (adjusted for BMI), MetS, gout, and Graves' disease;
To obtain GWAS summary data for these traits, we collected information on IVs from various sources for East Asian populations (Supplementary Table 1). BMI, uric acid, creatinine, gout, and Graves' disease IVs were obtained from disease characteristics released by Biobank Japan (BBJ) through the Open GWAS project of the Integrated Epidemiology Unit (IEU).14and IV for HDL-C, LDL-C, TC, TG, and T2D.15 These were obtained from the Asian Genetic Epidemiology Network (https://blog.nus.edu.sg/agen/). Furthermore, infusions according to the presence or absence of alcohol and coffee intake were extracted from the study of Mataba et al.16the sweetness IV was extracted from the study of Kawafune et al.17waist circumference and waist-to-hip ratio IV were extracted from the study of Wen et al.18, education level IV was extracted from the UK Biobank pan-ancestry genetic analysis performed at the Broad Institute (https://gwas.mrcieu.ac.uk/datasets/ukb-e-845_EAS/). The dataset (ukb-e-845_EAS) was collected within participants of East Asian descent. The IV of glycine was extracted from the study of Chang et al.19the IV of MetS was extracted from the study of Zhu et al.20gout infusion was extracted from the study of Nakayama et al.twenty one.
A series of quality control measures have been established to ensure the quality of instrumental single nucleotide polymorphisms (SNPs) of risk factors. First, the SNP is set to the conventional threshold (P< 5 × 10–8). We then removed SNPs with minor allele frequency (MAF) less than 0.01. Third, we only retained SNPs with long physical distance (window size = 10,000 kb) and low probability of linkage disequilibrium estimation (r2< 0.001) from East Asian samples from the 1000 Genomes Project. Fourth, the proportion of variance explained by SNPs (PVE) was calculated according to the following formula: PVE = 2 ×MAF ×(1-MAF) ×beta2twenty two. The strength of each SNP was measured by calculating the F statistic using the following formula: F =PVE× (N− 2)/(1− PVE) twenty two. If MAF was not available in the original study, another formula was used for calculation ( F= beta2/se2)twenty three.statistical power F> 10 was considered to indicate a strong association13.
Finally, we analyzed the 19 eligible risk factors separately. A summary of these genetic tools is provided in Supplementary Table 1. We also excluded candidate genetic tools that failed the MR Pleiotropy Residual Sum and Outliers (MR-PRESSO) test ( P< 0.05)twenty four. Ethical approval was obtained for the original study.
GWAS summary statistics for DSC
To ensure comparability in patient ancestry, genetic variants and EC (1,300 cases and 195,745 controls), GC (6,563 cases and 195,745 controls), CRC (7,062 cases and 195,745 controls), HCC (1,866 cases and 195,745 controls), BTC (339 cases and 195,745 controls), and PC (442 cases and 195,745 controls) were recruited from 12 medical institutions, mainly in Japan, during 2003. Retrieved from the Biobank BBJ, which contains data on approximately 200,000 participants. 200814,25. In the IEU open GWAS platform, the GWAS IDs corresponding to EC, GC, CRC, HCC, BTC, and PC are “bbj-a-117”, “bbj-a-119”, “bbj-a-107”, and “respectively. “bbj-a-158”, “bbj-a-92”, “bbj-a-140”. During the process of extracting exposure-related SNPs from the results, SNPs lacking relevant details in the results were excluded.26.
Overlapping participants in MR analysis
Duplication of participants in MR analysis can increase Type I error27, we attempted to select IVs from sources other than BBJ to avoid introducing bias due to sample overlap. However, genetic variations in certain risk factors such as BMI, uric acid, creatinine, and Graves' disease were only identified from BBJ. The potential bias introduced by duplicate sample of participants was calculated using the following formula: bias=βr/Fwhereβis the MR estimate,ris the sample overlap rate between the exposure and the outcome,FIt's averageFStatistics averaged across IVs27.
statistical analysis
I created a flowchart to perform MR step by step. First, we harmonized her GWAS data for risk factors and DSC with selected IVs that served as matching indicators. We then used the MR-PRESSO approach to detect pleiotropic outliers among the selected IVs and remove them before MR analysis. Third, we performed MR-Egger regression to test for horizontal pleiotropy.P>0.05 indicates no evidence of horizontal pleiotropy28. Additionally, this study aimed to confirm the association between the IVs included in the analysis and significant risk factors for cancer, such as smoking and family history of all malignant neoplasms. To perform this check, we employed an online tool known as PhenoScanner V2.29. If we found IVs associated with smoking or family history of malignant neoplasms, we excluded them and repeated the MR analysis.
After excluding pleiotropy, Cochran's Q test was used to detect heterogeneity between SNPs and different MR techniques were used to ensure directional consistency (i.e., MR-Egger regression, weighted median, weighted inverse variance). [IVW; random-effects models]weighted mode [WM]). A scatter plot was created to visualize the results. For sweet taste, waist-to-hip ratio, glycine, and MetS, only the Wald ratio (WR) method was used (≤2 SNPs). We calculated the odds ratio (OR) and corresponding 95% confidence interval (CI) for DSC for each 1 standard deviation (SD) increment for quantitative exposure or for each unit change in the log odds scale for binary exposure.twenty four. Finally, a leave-one-out sensitivity test was used to assess the robustness of the IVW estimates and detect potentially influential SNPs. Statistical power for MR was estimated using mRnd30 .
We applied a Bonferroni-corrected significance level.P< 2.63 × 10–3 (0.05/19), which indicates a strong association;PValue between 2.63 × 10–30.05 indicated a suggestive association. All analyzes were performed using the TwoSampleMR function of the R package (version 4.0.3).
Ethics approval and consent to participate
Our analysis is based on publicly available data approved by the relevant review board and requires no additional ethical approval or consent to participate.