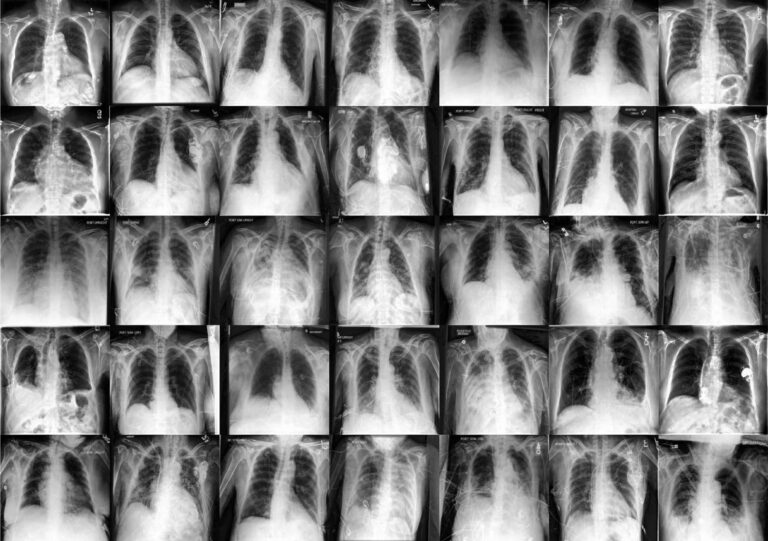
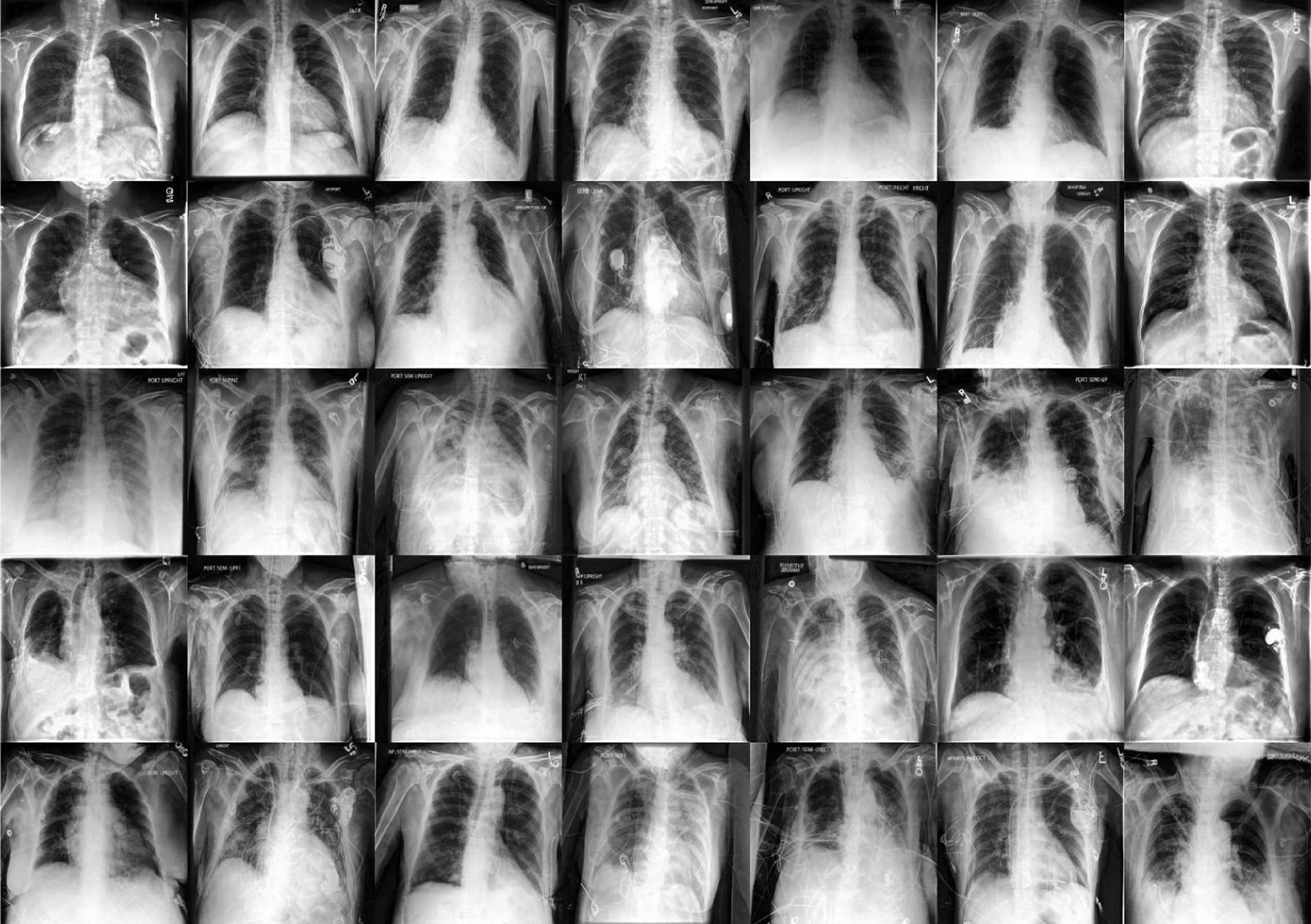
In a recent study published in the journal natural medicineResearchers used diffusion models for data augmentation to increase the robustness and fairness of medical machine learning (ML) models in three medical imaging contexts: histopathology, chest radiographs, and dermatology images. did.
 study: Generative models improve the fairness of medical classifiers under changing distributions
study: Generative models improve the fairness of medical classifiers under changing distributions
Domain generalization has become a major issue in the use of ML in healthcare settings, as data inconsistencies during model development and deployment can cause model performance to be worse than planned. Underrepresentation of certain groups or diseases is a typical problem that competent physicians struggle to solve due to the rarity of the disease or the availability of clinical knowledge. Few initiatives have achieved wide acceptance and significant impact on clinical outcomes, and “out-of-circulation” data remains a major hurdle to implementation.
About research
In this study, researchers used the diffusion model to examine medical imaging situations such as histology, chest radiographs, and dermatology photographs. They used these photos to enhance the reliability and fairness of their medical machine learning models. We also utilized unlabeled data to track the distribution of the data and supplement the real sample. In this project, we aimed to extend the training dataset in a manipulable and programmable way.
The researchers trained a generative model using labeled and unlabeled data. Labeled data can only be accessed from a single source domain, and additional unlabeled data can be accessed from any domain (within or outside the distribution). You can condition your model on diagnostic labels with or without properties (such as sensitive attribute labels or hospital IDs). The researchers anonymized the data before analysis. By conditioning the model on one or both qualities, you can now specify which synthetic samples are used to supplement the training set. They used a single conditioning vector to train low-resolution and upsampler generative models.
The team added synthetic images from generative modeling to training data obtained from the source domain before training the diagnostic model. They used a denoised diffuse probabilistic model (DDPM) to track fairness and diagnostic performance both inside and outside of delivery (OOD) and tested the strategy in several medical contexts. They defined within-distribution data as photographs from similar demographic and disease distributions that were acquired using one imaging technique as training data.
The researchers used two criteria to compare the baseline performance of the model with the proposed method. One set focused on diagnostic accuracy, such as top-1 type accuracy in histology identification and receiver operating curve (ROC-AUC) values for radiological evaluation, whereas the other One was focused on fairness. Expert dermatologists have found high-risk type hypersensitivity to be the most useful diagnostic tool.
The researchers used two large public radiology datasets, CheXpert and ChestX-ray, to create generative and diagnostic models for chest X-rays. After training on 201,055 chest radiographs, a dermatologist evaluated the model's ability to capture key features on his 488 composite photographs of normal and high-risk classes. They assessed image quality and provided diagnoses for up to three of approximately 20,000 common diseases.
result
This study shows that diffusion models can learn realistic extensions from data in a label-efficient manner, making them more resilient and statistically fair both within and outside of distribution. . Combining synthetic and real-time data can significantly improve diagnostic accuracy and reduce the equity gap between different qualities during delivery changes.
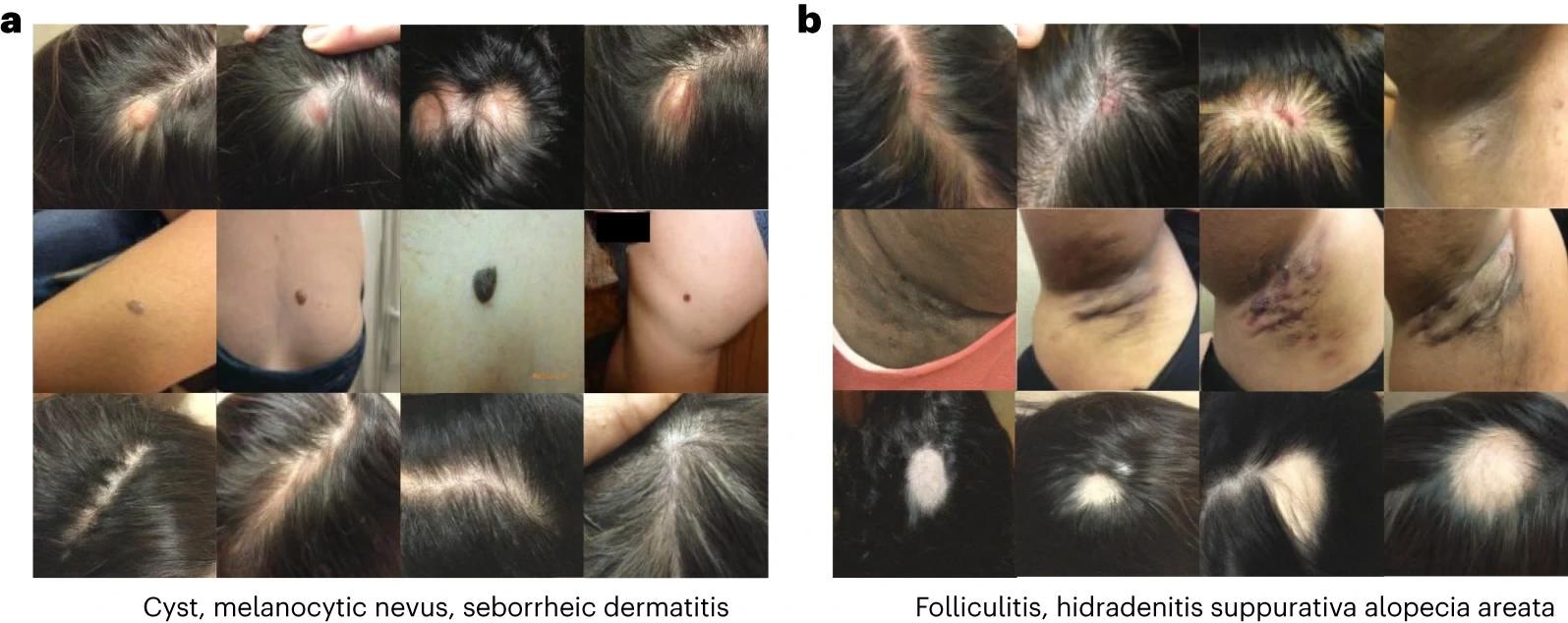 Image generated in a dermatology setting. Each row of the image corresponds to a different condition. begenerated images of cysts, melanocytic nevi, and seborrheic dermatitis. bgenerated images of folliculitis, hidradenitis, and alopecia areata.
Image generated in a dermatology setting. Each row of the image corresponds to a different condition. begenerated images of cysts, melanocytic nevi, and seborrheic dermatitis. bgenerated images of folliculitis, hidradenitis, and alopecia areata.
Although not a substitute for representative, high-quality data collection methods, clinicians can use unlabeled and labeled information to identify potential differences between under- and over-represented populations without penalty. can close harmful diagnostic accuracy gaps. The researchers found that using synthetic data outperformed the baseline in the distribution even in the presence of some skew, narrowing the equity gap between hospitals.
The color augmentation on the generated samples showed the best performance overall, with a relative improvement of 49% compared to the baseline modeling and compared to the model that underwent color augmentation training on the test hospital. An improvement of 3.2% was observed. This study showed that synthetic images significantly increased the average AUC of his five diseases, especially cardiac hypertrophy and OOD. The women's equity gap has narrowed by 45%, and the race equity gap has narrowed by 32%. Combining heuristic enhancements with synthetic database techniques such as “label conditioning” and “label and property conditioning” increases model sensitivity without sacrificing fairness and significantly improves OOD scenarios. .
Label and property adjustments improved high-risk diagnostic sensitivity by 27%, increased OOD by 63.5%, and narrowed the equity gap by 7.5x. The dermatology modality has created a realistic and standard picture that captures the characteristics of numerous diseases, including rare ones. The synthetic images also reduced spurious correlations and condensed representations, making her OOD correlations less generalizable and the model's dependence on underserved individuals.
This study shows that diffusion models have the potential to generate synthetic images useful for medical applications such as histology, radiology, and dermatology, while improving statistical fairness, balanced accuracy, and high-risk sensitivity. is shown. These synthetic samples produce realistic, standard images that expert physicians consider diagnostic. However, the researchers point out possible risks and limitations depending on the data produced, such as overconfidence in the AI system, limited insight, and recurrence of bias in the original training data.