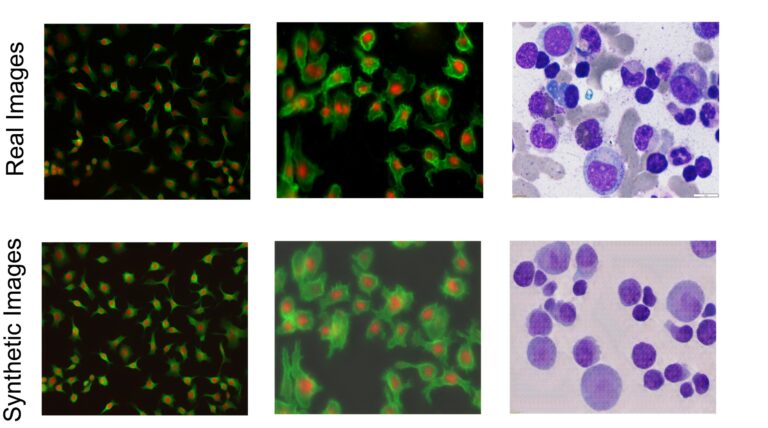
Examples of real images of different cell types and synthetic images generated by researchers' generative models.Credit: Ali Shariati et al.
× close
Examples of real images of different cell types and synthetic images generated by researchers' generative models.Credit: Ali Shariati et al.
Observing individual cells under a microscope reveals a variety of important cellular biological phenomena that are frequently involved in human diseases, but the processes that distinguish single cells from each other and their backgrounds are It is very time consuming and very suitable. For AI support.
AI models learn how to perform such tasks using sets of data annotated by humans, but the process of distinguishing cells from the background, called “single cell segmentation,” takes time. It also takes a lot of effort. As a result, the amount of annotated data available in the AI training set is limited. Researchers at the University of California, Santa Cruz are building microscopic image generation AI models to create realistic images of single cells, and to train AI models to better perform single-cell segmentation. We have developed a method to solve this problem by using it as “synthetic data”.
The new software is described in a new paper published in the journal iscience. The project was led by assistant professor of biomolecular engineering Ali Shariati and graduate student Abolfazul Zargari. This model, called cGAN-Seg, is available for free on GitHub.
“The images obtained from our model are ready to be used to train the segmentation model,” Shariati said. “In a sense, we are doing microscopy without a microscope, in that we can produce images that are very close to images of real cells in terms of morphological details of single cells. The beauty is when the cells appear. The model images are already annotated and labeled, which shows many similarities with the real images. , can generate new scenarios that the model did not recognize during training.
Images of individual cells viewed under a microscope can help scientists learn about cell behavior and dynamics over time, improve disease detection, and discover new drugs. Details inside cells, such as texture, can help researchers answer important questions such as whether a cell is cancerous.
However, manually finding and labeling cell boundaries from the background is very difficult, especially in tissue samples where there are many cells in the image. It can take several days for researchers to manually perform cell segmentation on just 100 microscopic images.
Deep learning can speed up this process, but an initial data set of annotated images is required to train the model. To train an accurate deep learning model, you need at least a few thousand images as a baseline. Even if a researcher were able to find and annotate 1,000 images, those images may not contain the changes in features that appear under different experimental conditions.
Examples of cell images before and after segmentation, a process that allows researchers to distinguish single cells from each other and their background.Credit: Ali Shariati et al.
× close
Examples of cell images before and after segmentation, a process that allows researchers to distinguish single cells from each other and their background.Credit: Ali Shariati et al.
“We want to show that deep learning models work across a variety of samples with different cell types and image quality,” Zargari says. “For example, if you train a model using high-quality images, it won't be able to segment low-quality cell images. It's rare to find such a good dataset in microscopy.”
To address this issue, researchers created an image-to-image generative AI model. This model takes a limited set of annotated and labeled cellular images and introduces more complex and diverse subcellular features and structures to generate more images and create a diverse “synthetic” set. Masu. image. In particular, it can generate annotated images containing a high density of cells that are particularly difficult to annotate manually and are particularly relevant for the study of tissues. This technology works to process and generate images for a variety of imaging techniques, including images of different cell types and images taken using fluorescence and histological stains.
Zargari, who led the development of the generative model, employed a commonly used AI algorithm called a “cycle generative adversarial network” to create realistic images. The generative model is powered by so-called “extensions” and “style injection networks,” which allow the generator to create a wide variety of high-quality synthetic images that show different possibilities of how the cells could look. Helpful. To the researchers' knowledge, this is the first time that style injection techniques have been used in this context.
This different set of synthetic images created by the generator is then used to train a model that accurately performs cell segmentation on new real images taken during the experiment.
“Using a limited dataset, you can train a good generative model. That generative model can then be used to generate a more diverse and larger set of annotated synthetic images. Using the generated synthetic images That's the main idea.
The researchers compared the results of the model using synthetic training data to traditional methods of training AI to perform cell segmentation across different cell types. They found that their model produced significantly improved segmentation compared to models trained using traditional limited training data. This confirmed to the researchers that providing a more diverse dataset while training a segmentation model improves performance.
These enhanced segmentation capabilities allow researchers to better detect cells and study variations between individual cells, especially stem cells. In the future, the researchers hope to use the technology they developed to go beyond still images and produce videos to determine which factors influence a cell's early lifespan and predict its future. I would like to help.
“We are generating composite images that can also be turned into time-lapse videos, where we can generate unseen futures of cells,” Shariati said. “We want to see if this allows us to predict the future state of cells, such as whether they will grow, migrate, differentiate, or divide.”
For more information:
Abolfazl Zargari et al., Enhanced Cell Segmentation with Restricted Training Datasets Using Cycle Generative Adversarial Networks, iscience (2024). DOI: 10.1016/j.isci.2024.109740
Magazine information:
iscience